SnapGene Version 4.1.6
SnapGene 4.1.6 was released on February 21, 2018.
Overview
This version includes additional fixes and enhancements.
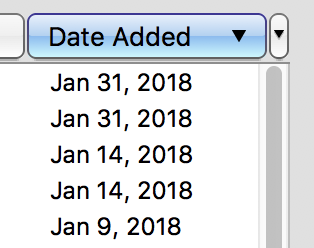
Sort a Collection List Bidirectionally
Sort the files in a collection in either order according to date or code number.
New Functionality
- Added the ability to press Alt/Opt while mousing over a sequence trace to see all four peak heights.
(Requested by Teisha Rowland) - Provided additional TaKaRa DNA markers.
(Requested by fky19842004)
Enhancements
- Enabled bidirectional sorting of a collection by Code Number, Date Added, or Date Created by clicking the column buttons.
- Made horizontal scrolling easier to discover by setting ∞ as the default fixed line width.
Fixes
- Fixed an issue on Windows where automatically launching the application after running the installer resulted in the settings being stored in the wrong location and administrator file ownership and permissions being used when saving.
(Reported by Clay Fuqua) - Improved the output when exporting and printing agarose gels, especially while using high-DPI displays.
- Corrected an erroneous message that was shown when exporting common features.
(Reported by Edward Wallace) - Ensured that gray sequence colors are visible after saving and re-opening a file.
(Reported by Genevieve Tauxe) - Fixed a regression to ensure that gaps in aligned sequences are always visibly represented.
(Reported by Dustin Rubinstein) - Enhanced the importer for CLC Bio files.
(Reported by Rabea Wagner) - Fixed an issue with opening certain DNA Strider files.
(Reported by Michael Nassal) - Improved the import of primers from SeqBuilder files.
(Reported by Lakxmi Subramanian) - Ensured that error messages in the primer dialogs are completely visible.
- Enhanced the flexibility of controls for adding and importing primers copied to the clipboard.
(Requested by Seth Goldman) - Prevented a slowdown and potential data omission that could occur when using the same open file for multiple inserts in a manipulation dialog.
(Reported by Seth Goldman) - Ensured that modifications to open documents are reflected in manipulation dialogs that already have those documents loaded.
- Prevented duplicate forward slashes from getting into the recent collections list.
- Improved the default location for saving sequences in an imported archive.
- Prevented unwanted data from being retrieved from an imported sequence record when including extra features added by NCBI.
- Blocked accidental toggling of the "File Name" column in a collection list.










