SnapGene Version 4.2.0
SnapGene 4.2.0 was released on July 17, 2018.
Overview
Note: SnapGene 4.2 requires macOS 10.10 or later.
Version 4.2 adds an interface for multiple alignment, together with improved tools for displaying and exchanging data.
Multiple Protein and DNA Alignments
Align protein or DNA sequences using any combination of Clustal Omega, MAFFT, MUSCLE, or T-Coffee. A versatile interface provides options for rendering and editing an alignment.

Improved Display of Related Features
pre-mRNA features such as exons are automatically grouped for display in the same row of a map.

Tabbed Interface for macOS
With macOS 10.13 and later, sequence windows can now be organized with tabs.

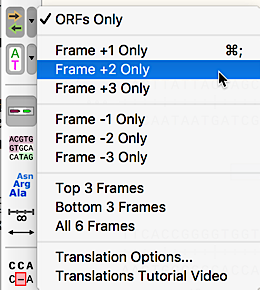
Choice of Any Reading Frame
In Sequence view, choose any of the six reading frames for a full-sequence translation.
File Exchange with the LabArchives ELN
Exchange files between SnapGene and the LabArchives Electronic Lab Notebook (ELN).

New Functionality
- Added multiple alignment tools for DNA and protein sequences.
(Requested by hundreds of customers) - Implemented tabbed window support for macOS.
(Requested by Kasey and others) - Enabled import of a database in CSV format into a collection.
(Requested by James Burchfield) - Enabled display in Sequence view of any of the six reading frames.
(Requested by Assunta Diodato, Matthias Goerlach, Paul Shapiro, and others) - Added file exchange tools for the LabArchives ELN.
- Allowed sequence traces to be exported to FASTA and plain text formats.
- Added Biotium MW markers.
(Requested by Hussein Abkallo) - Added "φ29 – HindIII" MW Marker.
(Requested by Manuel Miras Marín) - Added "λ DNA – EcoT14I" MW Marker.
(Requested by Nori) - Added Enzynomics MW markers.
(Requested by the Department of Parasitology and Tropical Medicine) - Restored original "1 Kb Plus DNA Ladder" from Thermo Fisher (Invitrogen).
(Requested by Martin Schmidt, Aminu Jahun, Chris McDermott-Roe and Shih-wei Chen) - Facilitated unattended installations by adding support for accepting the EULA and registering and unregistering via the command line interface.
Enhancements
- Improved the placement of features in Map view so that certain types of related features are in the same row.
- Made the selection behavior more consistent and predictable in Map view, especially in circular maps.
(Requested by Thomas Seine and Elisabeth) - Increased the number of allowed fragments to ten when inserting or assembling.
(Requested by Álvaro Muñoz-López) - Added the "Set Translation Numbering..." command to relevant context menus.
(Requested by Karl Brune) - Enhanced the mouse indicator to indicate the offset within the selection when mousing over DNA bases in Sequence view.
(Requested by Steven Erickson) - Improved the drag selection behavior in circular maps.
(Requested by Elisabeth) - Improved the Find tool to show the closest match to the currently displayed portion of the sequence instead of the first match in the entire sequence.
(Requested by Adrian Ross) - Made extensive speedups and other optimizations.
- Enabled the NCBI importer to accept UniProt accession numbers.
- Moved options related to files to a separate tab in the Preferences dialog, and added new options for working with files.
- Added the option to hide the Order button, and added a "How Ordering Works" command.
- Added a "Note About Tm Values" command that explains how Tm values are calculated.
- Enhanced the "Format Tips" window to explain use of the COMPLEMENT operator when importing from NCBI.
- Improved inserting codons and other content into a primer sequence by placing the cursor as one would intuitively expect.
- Improved the default view tab shown when using "New File from Selection."
- Migrated "Add Custom Codon Usage Table" to a dedicated button to improve discoverability.
- Added "Choose Alternative Codons..." to the translated feature context menus.
- Enhanced the menus in the top toolbar to provide commands relevant to collections, and added "Open File" to the Open menu.
- Streamlined the controls in Preferences for specifying if 1- or 3-letter codes should be shown by default, by offering to mirror changes for DNA and protein sequences.
- Added a link to "Preferences" to the launch dialog on Windows.
- Updated the Restriction Enzymes and Common Features databases.
- Updated the embedded vector sets for Gateway cloning and TA and GC cloning.
- Made various color, textual, visual alignment and rendering enhancements.
Fixes
- Ensured that "Find protein sequence" will return matches to feature translations that span multiple features.
(Requested by John Leonard, Patrick McGrath, and Omar Bazirgan) - Improved the display of selected amino acids at the ends of rows in Sequence view.
(Reported by Dan Strongin) - Fixed an issue with displaying feature translations when viewing primer binding sites at or near the numerical origin.
(Reported by Lagnajeet Pradhan) - Enabled use of the command line interface on Linux without an X server.
(Requested by Novozymes) - Improved stability when using SnapGene on Linux via VNC.
(Reported by Tom Chappell) - Fixed an issue with highlighting matches and showing selections in sequence traces.
(Reported by Moin) - Ensured that the "DNA Calculations" window always shows information about the active selection.
(Reported by Scott James) - Ensured consistent letter spacing in Sequence view on Windows when using multiple screens.
(Reported by Gian Mario Dore) - Preserved the file location in the folder hierarchy when renaming a file in a collection on Windows.
(Reported by Carles Alvarez ) - Preserved the original numbering of protein sequences when using "New File from Selection".
(Reported by Karl Brune) - Fixed an issue with saving a file into the proper collection folder.
(Reported by Carles Alvarez) - Fixed an issue with using files stored on an SMB network share via the Gnome Virtual File System (GVFS) protocol on Linux.
(Reported by Kai Zhou) - Corrected a stability issue when renaming folders in collections on macOS.
(Reported by Jorge) - Fixed an issue with embedding and resurrecting ancestors to protein sequences, and with undoing protein sequence modifications.
(Reported by Karl Brune) - Ensured that files saved to collections always appear in Folder and List views.
(Reported by Carles Alvarez) - Sped up addition, removal, and editing of custom common features on Windows.
(Reported by Wulf Dirk) - Improved the unified toolbar appearance on macOS High Sierra.
- Provided real-time feedback so that customizing the top toolbar immediately updates all toolbars to show the new options.
- Ensured that right clicking does not modify the selection in a list in the Restriction Enzymes and Simulate Agarose Gel dialogs.
- Enhanced History view so that if the display is limited to one level, history colors are shown for the last logged ancestor and operation.
- Improved the display of features in aligned sequences that do not exactly match the reference sequence.
- Improved decoding of the comment line from FASTA files.
- Improved the rendering of features.
- Improved the selection behavior in lists.
- Fixed an issue where the second segment in a multi-segment feature was selected automatically when using the Edit Feature command.
- Improved the decoding of non-UTF-8-encoded GenBank files that contain Greek symbols and other characters outside the normal alphabet.
- Ensured that file dialogs persist appropriately on macOS.
- Disabled the "Export DNA" command in the Simulate Agarose Gel dialog and other dialogs where it was nonfunctional.
- Improved the display of a primer description when space is limited and the primer has multiple binding sites.
- Improved the rendering of overlapping selections and yellow "Find" highlighting in Sequence view.
- Improved the reliability of detecting the topology in some GenBank files.
- Added logic to preserve the creation date from GenBank files.
- Simplified the oligo name controls in the Anneal Oligos dialog.
- Added licensing information to the Linux .deb package.
- Fixed an issue with included plasmids with single quotes in their names in the Linux .rpm package.
- Improved the import of primers from FASTA-formatted lists that are RTF encoded or compressed.
- Configured tooltips in protein sequences to persist rather than disappearing after a brief delay.
- Ensured that the source menu in a cloning dialog will list the active template file from a collection even if the collection is closed.
- Fixed a bug where source menus might not always select recently browsed files with very long names.
- Ensured that clicking Escape always hides the Find controls when viewing a sequence trace.
- Ensured that if the clipboard contains a valid protein sequence but lacks A's, C's, G's and T's, the protein window will still allow pasting.
- Prevented the "Import NCBI Sequences" command from hanging if no terminator symbol (//) is included at the end of the downloaded content.
- Ensured that exact searches for queries that include U will produce correct matches on the bottom strand of DNA sequences.
- Ensured that "Undo" will revert the modification date shown in the Description panel.
- Fixed an issue with ordering an edited but unsaved DNA sequence from VectorBuilder.
- Corrected links from the interface to open the relevant tab in Preferences.
- Fixed an issue where after clearing the Sequence Author, the previous value returned unexpectedly when modifying another field in the Description Panel.
- Improved the algorithms for displaying, updating, and copying cleavage arrows that are located at an end of a feature.
- Improved the display of feature codon selections in Sequence view.
- Fixed the listing in the Window menu of open collections with selected files.
- Improved the reliability of keyboard shortcuts for commands in cascading menus on macOS.
- Provided automatic selection of the "From" value in "Edit → Select Range" when no selection is present.
- Improved the conversion of UniProt sequences.
- Prevented the application from hanging when switching from a placeholder file to a non-empty sequence in a collection.
- Disabled "View → Edit Map Labels" when no sequence is being viewed.
- Improved batch conversion of files.
- Improved the identification of CPU and related information on Linux.
- Increased stability when exporting a selection.
- Fixed a stability issue with opening compressed sequence trace files.
- Fixed an issue with saving sequence traces using the ZTR1, ZTR2, and ZTR3 file formats.










