What is In-Fusion Cloning?
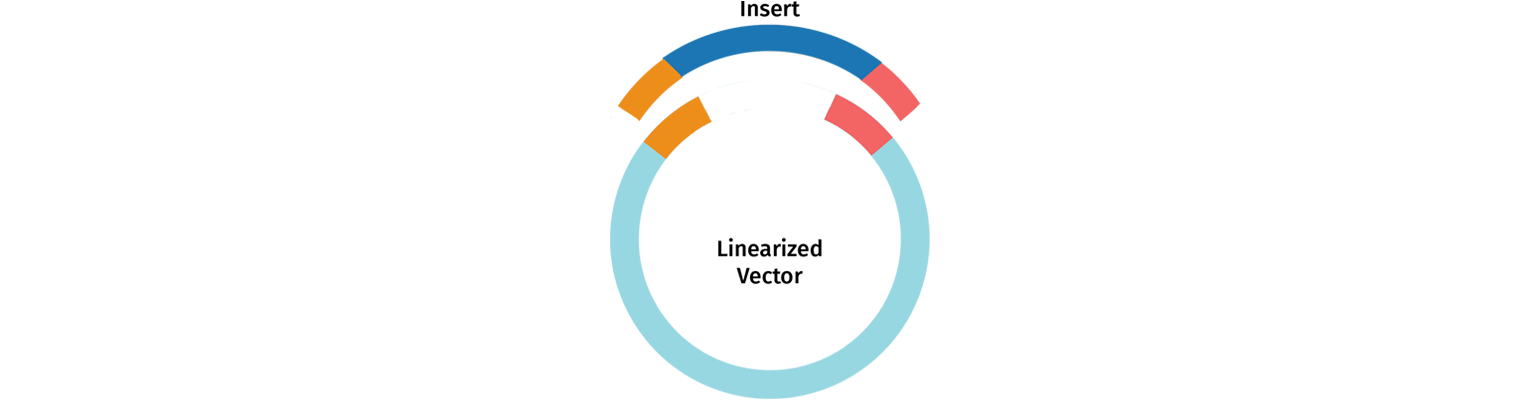
In-Fusion cloning is a remarkably versatile method developed by Takara Biosciences for creating seamless gene fusions. For the In-Fusion reaction, a linearized vector is mixed with one or more PCR products with overlapping ends. Like other PCR-based advanced cloning techniques, In-Fusion Cloning allows you to adjoin your fragments of interest with appropriately designed primers directly. This bypasses one of the major limitations of restriction enzyme cloning, making In-Fusion a sequence-independent and seamless cloning technique.
How does In-Fusion Cloning work?
In-Fusion Cloning utilizes the DNA polymerase from Vaccinia Virus. Like many DNA polymerases, while the preferred reaction is the addition of nucleotides, this polymerase contains 3’ → 5’ exonuclease activity that is part of its proofreading mechanism. In the absence of dNTPs, and when provided a substrate of double-stranded DNA, the exonuclease will chew back each 3’ end, creating asymmetrical overhangs. Overhangs can be engineered into cloning experiments by including the appropriate sequences in PCR primers.
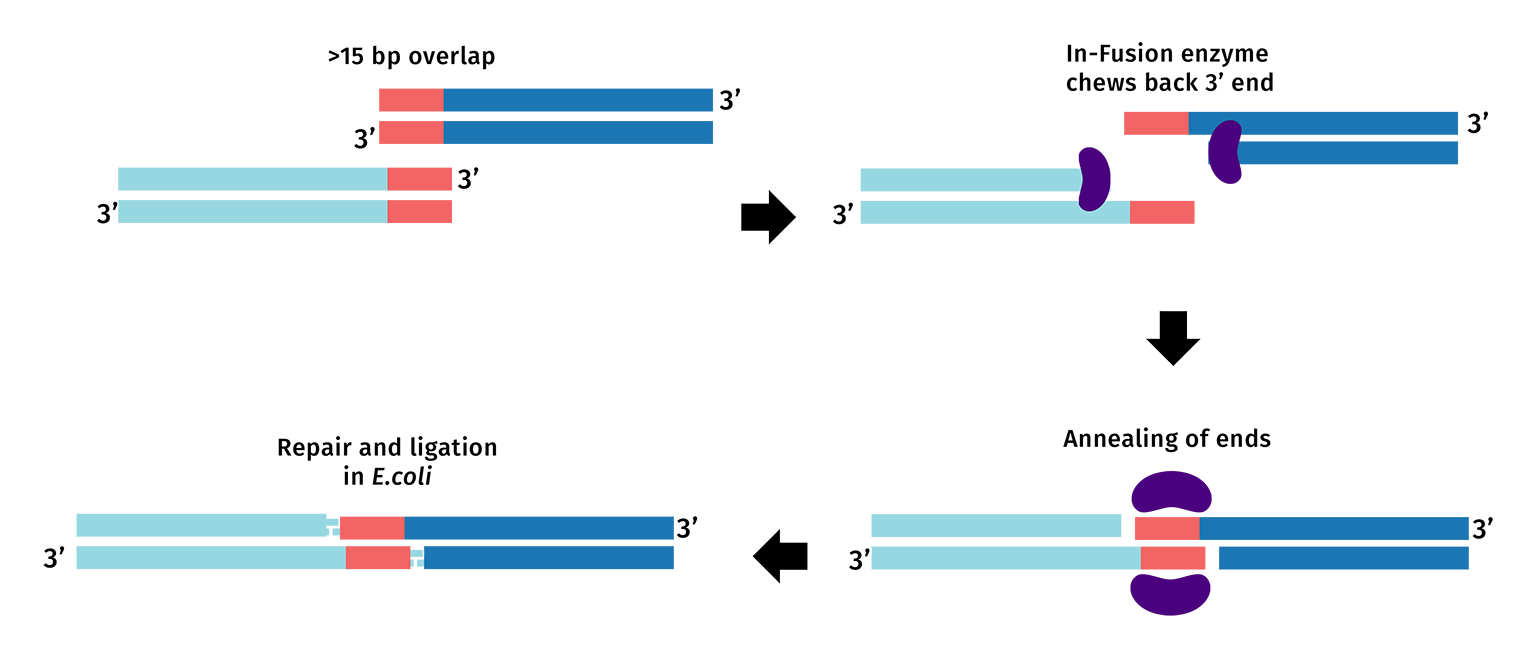
PCR primers are designed with an added 5’ tail to create 15 to 20 base pair overlaps between the fragments you intend to clone. During the In-Fusion reaction, the exonuclease activity from the polymerase chews back the 3’ ends, exposing the overlapping nucleotides.
The fragments are ligated in vivo after transformation into E. coli.
► Learn more about PCR cloning
Designing an In-Fusion Cloning Experiment
In-Fusion cloning allows you to add any insert into any vector at any site making it an extremely versatile cloning method.
The three main steps when designing your In-Fusion experiment are:
- Linearizing your vector at the insertion point of interest
- Designing PCR primers to amplify your insert of interest and add the necessary tail for annealing to your vector
- Performing the In-Fusion reaction and validating the fusion points
Linearizing the Vector
The first step in In-Fusion cloning is to linearize your vector of interest at the insertion site. Any vector suitable for the downstream experimental use can be used in the In-Fusion reaction.
The two main methods of vector linearization are restriction enzyme digestion and inverse PCR.
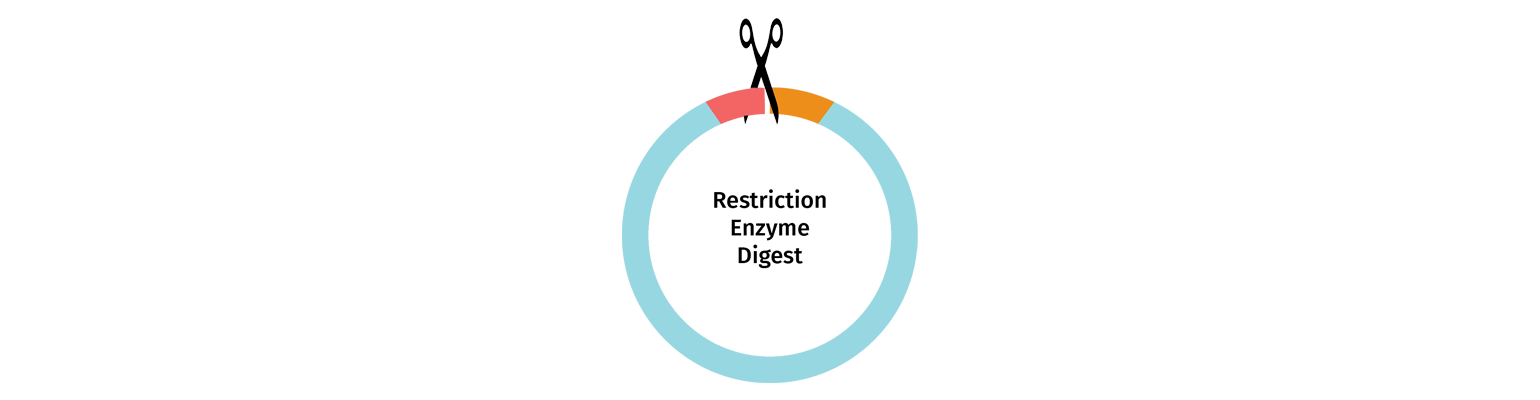
Restriction Enzyme Digestion
Vectors may be linearized using restriction enzyme digestion with either one or two enzymes selected to cut at the insertion point.
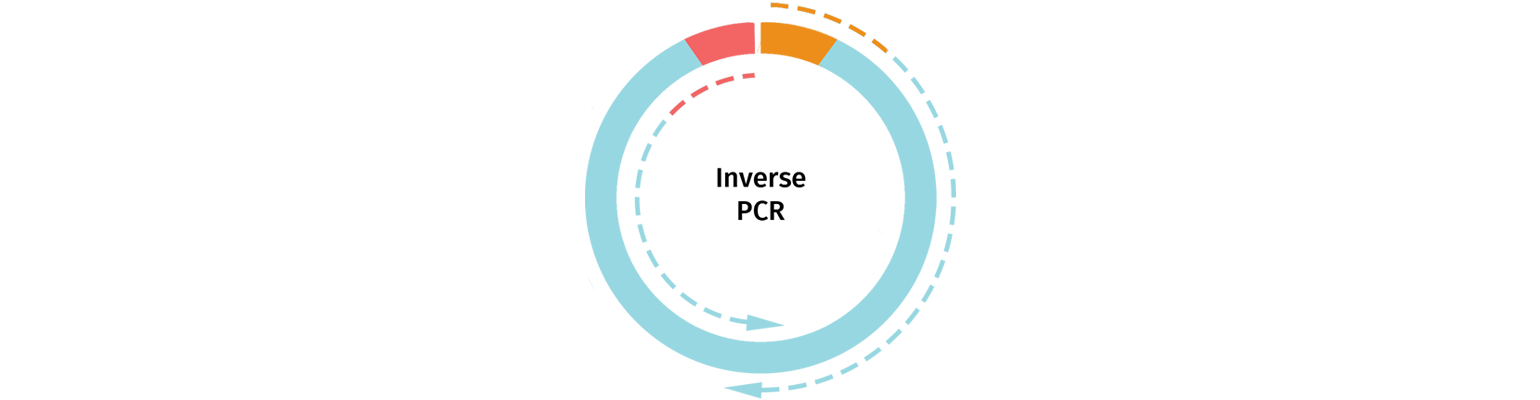
Inverse PCR
Linearized vectors may also be generated using Inverse PCR. Primers are selected that will amplify the backbone of your chosen cloning vector, removing the section of the vector that will be replaced by the insert.
Designing Primers for Single Insert Cloning
Single insert In-Fusion cloning requires a PCR reaction that amplifies the insert of interest and adds the necessary nucleotides for annealing to the vector. This is achieved by designing PCR primers containing- the template-specific portion and the vector-specific tail.
The template-specific primers are designed to amplify the insert of interest and are oriented with the 3' end of the primer facing into your target sequence. The design of these primers should be based on standard primer design guidelines.
The tail of each primer introduces a sequence that will allow assembly of the fragments after treatment with In-Fusion. The tails should be 15 nucleotides in length and complement the sequence of the vector end.

Designing Primers for Multi-Insert Cloning
Multi-insert In-Fusion cloning follows the same logic as single insert cloning, with primers designed to amplify the insert of interest and to create overlapping tails.
The template-specific part of the primer is designed to amplify each insert of interest with standard primer design.
The tail of each primer needs to be designed to overlap the tail of the adjacent insert or vector depending on the desired position. Longer tails (20 nt or greater) increase cloning efficiency when performing multi-fragment In-Fusion.
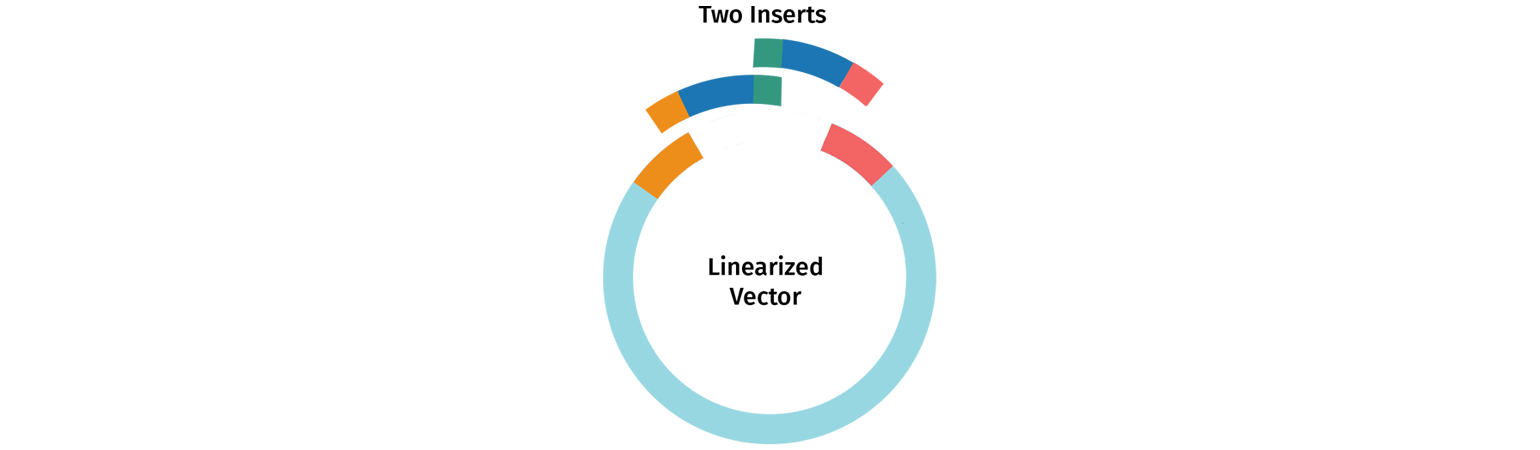
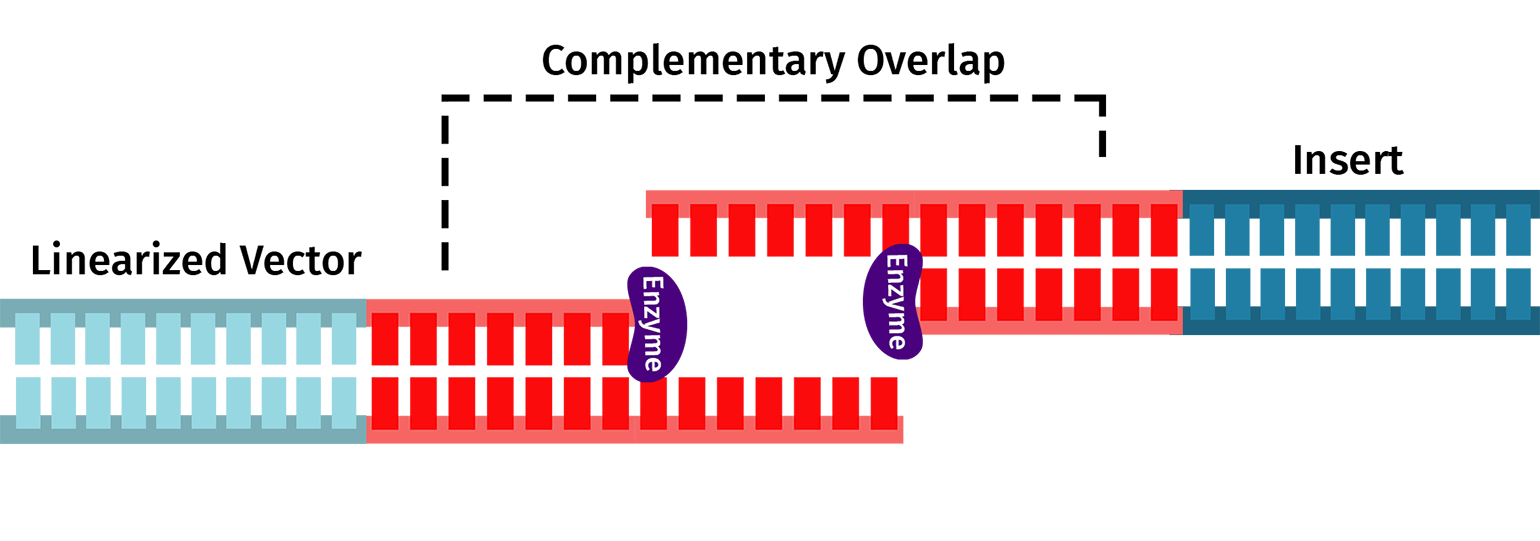
The In-Fusion Cloning Reaction
After amplification and clean-up, combine your purified insert and vector with the In-Fusion reagent. Incubate your fragments with the In-Fusion reagent for 15 minutes at 50ºC.
The In-Fusion reagent will chew back the nucleotides from the 3' ends of the linearized DNA allowing the complementary overlaps to join and anneal. You can then transform the mixture into E coli, where E coli's DNA repair machinery will create an intact plasmid.
Top Tips for In-Fusion Cloning
1. Primer Design tips
If you plan on using In-Fusion to clone various inserts into the same vector, you can design primer tails to be used on all pairs of primers and create one master vector preparation to use for all your cloning reactions.
2. Optimize the Length of Fragment Overlaps
- Single Fragment Insertion:
Overlapping regions of 12-15 base pairs are sufficient. - Multi-Fragment Insertions:
Overlapping regions 20 base pairs or more increases cloning efficiency.
3. Secondary Structure in Primer Tails
As Primer tails increase in length, pay attention to the stability of the secondary structures that the tails can form. Due to their length, many primer tails do not form stable secondary structures during PCR, where the reaction temperature ranges from 60°C to 75°C. In contrast, the In-Fusion exonuclease reaction is performed at 50°C, allowing some secondary structures to interfere with the exonuclease.
4. Quantitation
Quantitation of your inserts is important but not essential for In-Fusion Cloning. Takara Biosciences recommends the following molar ratios:
- Single Fragment Insertion
If your fragment and vector are similar in size, a molar ratio of 1:1 to 2:1 insert to vector is recommended.
As your insert fragment decreases in length (150 base pairs to 500 base pairs), you may wish to increase the molar ratio to between 3:1 and 5:1 insert to vector. Smaller inserts may require even greater molar ratios. - Multi-Fragment Insertion
When incorporating multiple fragment inserts into one vector, start with a molar ratio of 2 molar units for each insert for each single molar unit of vector, i.e., 2:2:1 for two inserts into one vector.
5. Reaction Kinetics
The primary catalytic function of the In-Fusion reagent is its exonuclease activity. The goal is to expose the engineered overlaps between your fragments. Adhere to the recommended fifteen-minute incubation.
Excessive nuclease activity will create exposed single-stranded ends that are longer than desired. This can lead to reduced reaction efficiency because the longer strands are unstable during DNA repair, and/or the longer single-stranded DNA may be subject to degradation.
In-Fusion Cloning: Pros & Cons
In-Fusion Pros
As one of several synthetic biology assembly techniques, In-Fusion is a sequence-independent and seamless cloning technique.
- The use of PCR means that reliance on conveniently located restriction enzymes is eliminated.
- The use of non-restriction and ligation cloning technology allows the direct fusion of desired fragments.
- Finally, the technique is relatively fast.
In-Fusion Cons
In-Fusion has a few limitations.
- The In-Fusion enzyme blend is proprietary to Takara Bio-sciences, entailing associated reagent costs.
- Because the In-Fusion reaction does not result in covalently joined DNA molecules, it is important to stick to recommended times for all steps in the procedure.
- Overlapping regions with highly stable secondary structures may resist exo-nuclease activity. If homologous overlaps are not exposed, the In-Fusion reaction will not succeed.
- Like any PCR-based cloning technique, sequence verification of your final clone is absolutely required.
In-Fusion Cloning in SnapGene
SnapGene allows you to accurately design and simulate In-Fusion Cloning reactions and supports the design of multi-fragment cloning products with as many as 100 inserts.
SnapGene’s Actions menu provides access to the In-Fusion Cloning tool. To plan an In-Fusion reaction, just select the DNA fragments you wish to fuse, and SnapGene will choose suitable primers.
Learn More About Cloning





